Tree operations¶
The objective of this notebook is to learn how to perform tree operations, such as subsetting and attachment.
Importing modules and basic settings¶
[1]:
import warnings
warnings.filterwarnings("ignore")
import scanpy as sc
import scFates as scf
sc.set_figure_params()
[2]:
adata = sc.read("data/empty_tree.h5ad",backup_url="https://github.com/LouisFaure/scFates_notebooks/raw/main/data/empty_tree.h5ad")
[3]:
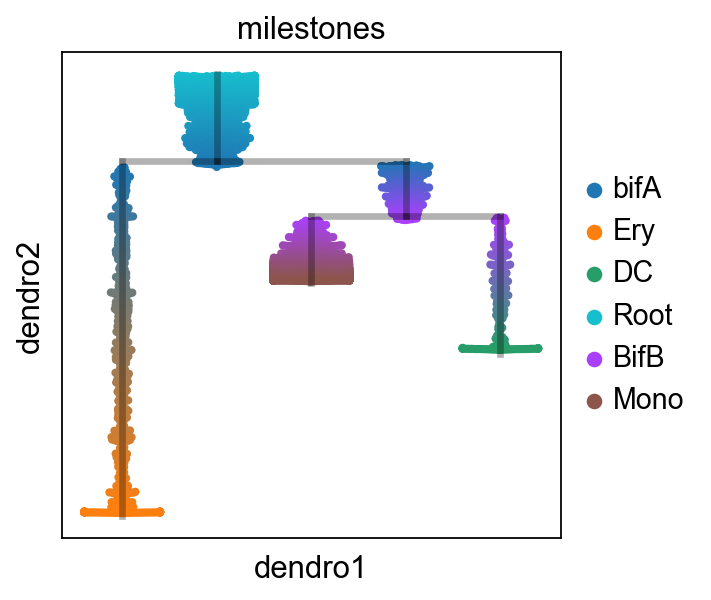
scf.pl.trajectory(adata,basis="draw_graph_fa",color_cells="milestones")

[4]:
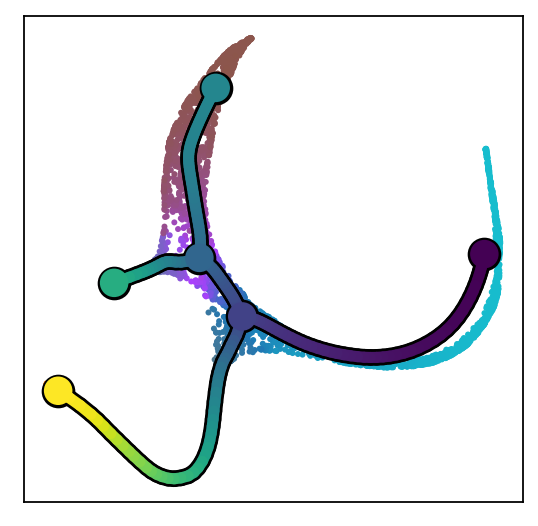
scf.pl.dendrogram(adata,color_milestones=True,color="milestones")
Subsetting of tree (extraction mode)¶
Subtree¶
[5]:
adata_extracted=scf.tl.subset_tree(adata,root_milestone="bifA",milestones=["Mono","DC"],copy=True)
subsetting tree
node 33 selected as a root --> added
.uns['graph']['root'] selected root.
.uns['graph']['pp_info'] for each PP, its distance vs root and segment assignment.
.uns['graph']['pp_seg'] segments network information.
projecting cells onto the principal graph
finished (0:00:01) --> added
.obs['edge'] assigned edge.
.obs['t'] pseudotime value.
.obs['seg'] segment of the tree assigned.
.obs['milestones'] milestone assigned.
.uns['pseudotime_list'] list of cell projection from all mappings.
finished (0:00:00) --> tree extracted
--> added
.obs['old_milestones'], previous milestones from intial tree
[6]:
ax=sc.pl.scatter(adata,basis="draw_graph_fa",color="whitesmoke",show=False)
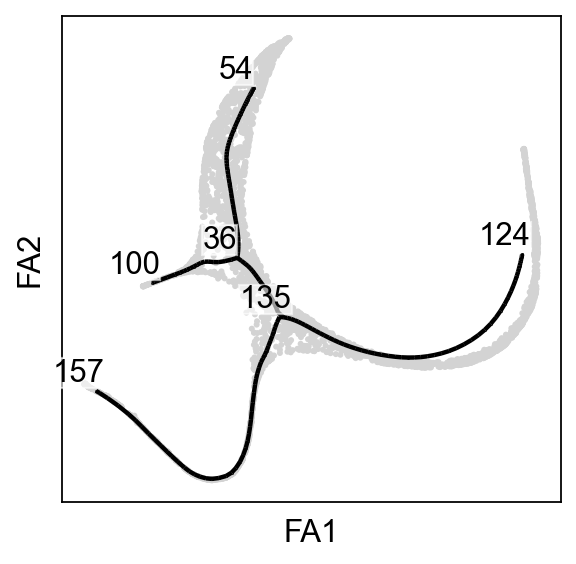
scf.pl.graph(adata_extracted,basis="draw_graph_fa",size_nodes=.1,ax=ax)
[7]:
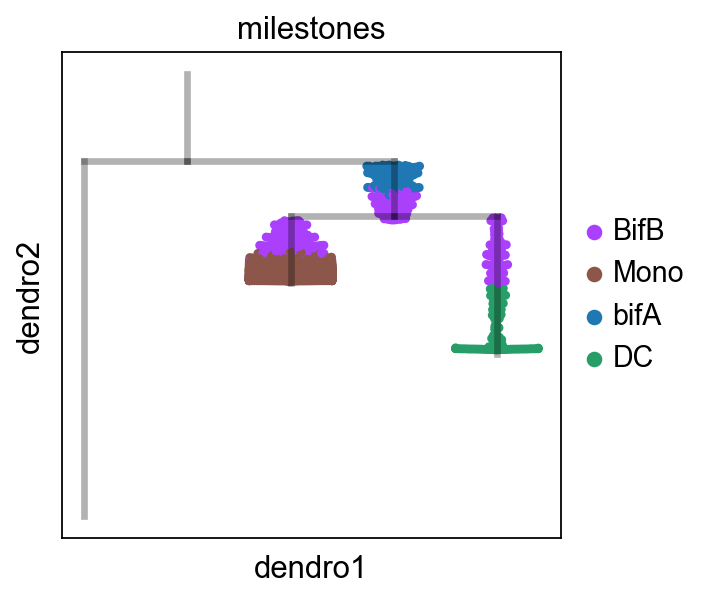
scf.pl.dendrogram(adata_extracted,color="milestones")
Single branch¶
[8]:
adata_extracted=scf.tl.subset_tree(adata,root_milestone="bifA",milestones=["Ery"],copy=True)
subsetting tree
node 42 selected as a root --> added
.uns['graph']['root'] selected root.
.uns['graph']['pp_info'] for each PP, its distance vs root and segment assignment.
.uns['graph']['pp_seg'] segments network information.
projecting cells onto the principal graph
finished (0:00:01) --> added
.obs['edge'] assigned edge.
.obs['t'] pseudotime value.
.obs['seg'] segment of the tree assigned.
.obs['milestones'] milestone assigned.
.uns['pseudotime_list'] list of cell projection from all mappings.
finished (0:00:00) --> tree extracted
--> added
.obs['old_milestones'], previous milestones from intial tree
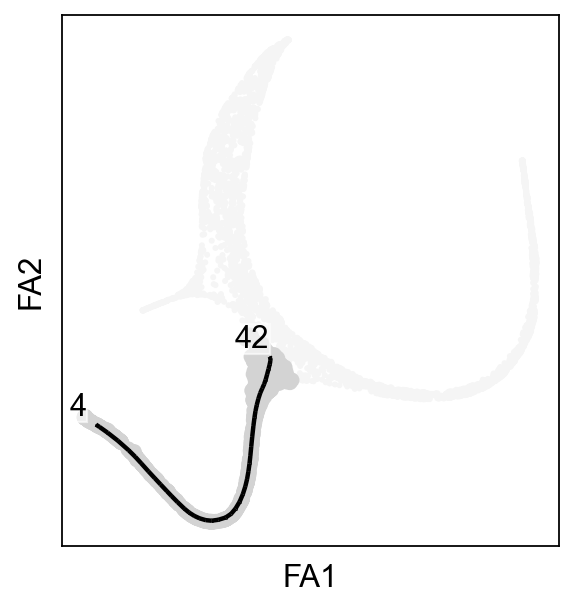
[9]:
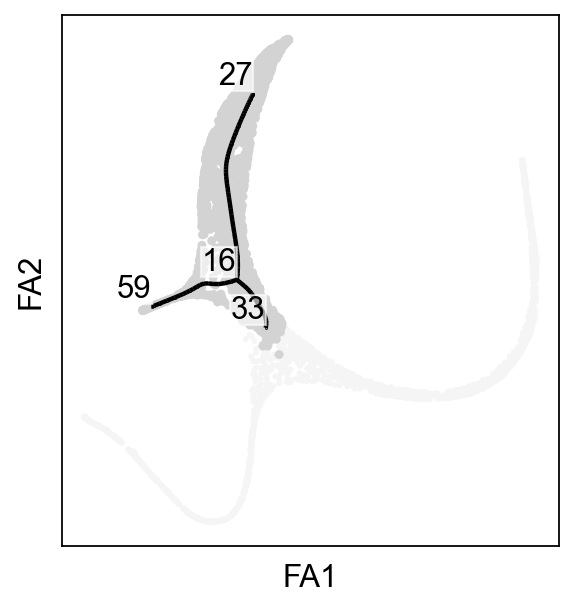
ax=sc.pl.scatter(adata,basis="draw_graph_fa",color="whitesmoke",show=False)
scf.pl.graph(adata_extracted,basis="draw_graph_fa",size_nodes=.1,ax=ax)
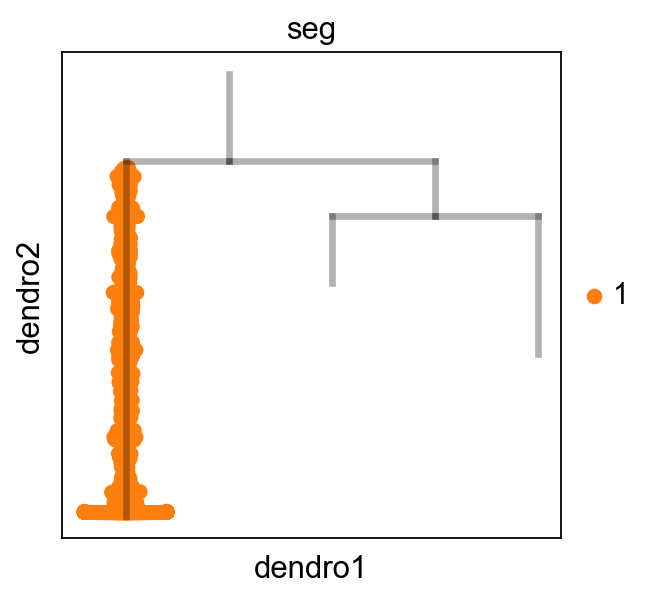
[10]:
scf.pl.dendrogram(adata_extracted,color="seg")
Subsetting of tree (substract mode)¶
[11]:
adata_subtracted=scf.tl.subset_tree(adata,root_milestone="bifA",milestones=["Ery"],
mode="substract",copy=True)
subsetting tree
node 124 selected as a root --> added
.uns['graph']['root'] selected root.
.uns['graph']['pp_info'] for each PP, its distance vs root and segment assignment.
.uns['graph']['pp_seg'] segments network information.
projecting cells onto the principal graph
finished (0:00:03) --> added
.obs['edge'] assigned edge.
.obs['t'] pseudotime value.
.obs['seg'] segment of the tree assigned.
.obs['milestones'] milestone assigned.
.uns['pseudotime_list'] list of cell projection from all mappings.
finished (0:00:00) --> tree subsetted
--> added
.obs['old_milestones'], previous milestones from intial tree
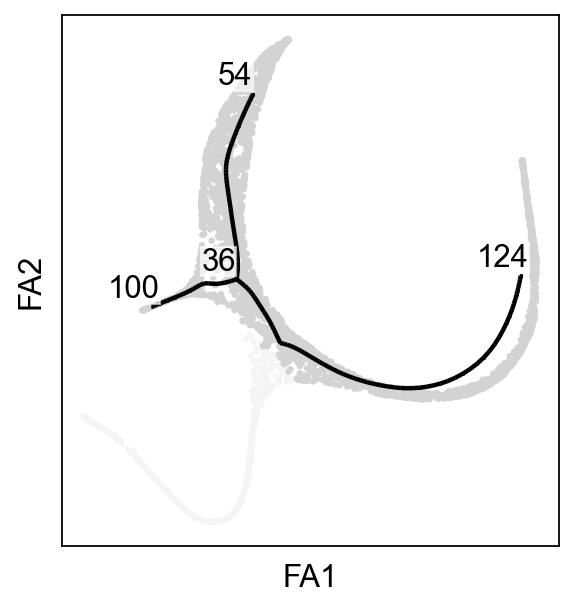
[12]:
ax=sc.pl.scatter(adata,basis="draw_graph_fa",color="whitesmoke",show=False)
scf.pl.graph(adata_subtracted,basis="draw_graph_fa",size_nodes=.1,ax=ax)
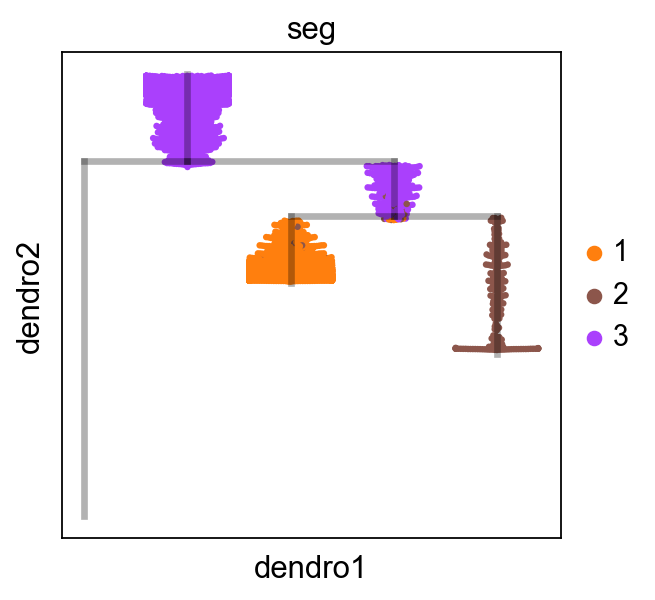
[13]:
scf.pl.dendrogram(adata_subtracted,color="seg")
Tree attachment¶
[14]:
adata_attached = scf.tl.attach_tree(adata_subtracted,adata_extracted)
attaching tree
merging
tree refitting
finished (0:00:02) --> datasets combined
[15]:
scf.pl.graph(adata_attached,basis="draw_graph_fa",size_nodes=.1)